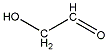
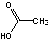
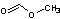Methods with standard basis sets
|
|
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
cc-pV(T+d)Z |
daug-cc-pVTZ |
| hartree fock |
HF |
98.6 a
33.3 b
0.0 c
28.0 d |
129.6 a
214.6 b
0.0 c
54.5 d |
129.6 a
214.6 b
0.0 c
54.5 d |
116.5 a
258.0 b
0.0 c
75.5 d |
b |
123.5 a
226.1 b
0.0 c
70.2 d |
118.8 a
228.4 b
0.0 c
71.5 d |
123.5 a
217.9 b
0.0 c
57.8 d |
120.9 a
230.4 b
0.0 c
70.9 d |
121.8 a
224.9 b
0.0 c
68.0 d |
116.6 a
0.0 c
67.7 d |
116.9 a
226.0 b
0.0 c
69.0 d |
121.5 a
232.0 b
0.0 c
71.4 d |
118.4 a
226.5 b
0.0 c
69.0 d |
112.6 a
229.2 b
0.0 c
74.6 d |
116.1 a
225.2 b
0.0 c
68.8 d |
118.4 a
0.0 c
69.0 d |
116.2 a
404.7 b
0.0 c
68.9 d |
| density functional |
LSDA |
|
NC |
NC |
NC |
NC |
NC |
NC |
NC |
NC |
NC |
|
|
NC |
NC |
NC |
|
|
|
| BLYP |
118.9 a
94.1 b
0.0 c
5.8 d |
124.7 a
223.9 b
0.0 c
37.0 d |
124.7 a
223.9 b
0.0 c
37.0 d |
116.7 a
246.6 b
0.0 c
52.6 d |
115.7 a
213.9 b
0.0 c
44.2 d |
115.3 a
224.6 b
0.0 c
54.5 d |
111.8 a
231.8 b
0.0 c
61.4 d |
a
b
d |
117.7 a
242.9 b
0.0 c
61.5 d |
112.9 a
223.4 b
0.0 c
53.6 d |
|
|
114.0 a
230.9 b
0.0 c
58.3 d |
113.6 a
238.0 b
0.0 c |
0.0 c
65.6 d |
|
113.6 a
0.0 c |
|
| B1B95 |
118.7 a
79.3 b
0.0 c
18.7 d |
140.1 a
226.8 b
0.0 c
50.8 d |
140.1 a
226.8 b
0.0 c
50.8 d |
132.8 a
258.7 b
0.0 c
66.8 d |
121.2 a
201.0 b
0.0 c
55.2 d |
130.9 a
222.8 b
0.0 c
65.9 d |
127.9 a
228.5 b
0.0 c
70.3 d |
134.6 a
222.3 b
0.0 c
57.5 d |
131.9 a
235.4 b
0.0 c
70.5 d |
220.1 b
0.0 c
64.6 d |
|
|
128.5 a
227.3 b
0.0 c
69.0 d |
117.0 a
217.9 b
0.0 c
69.9 d |
217.7 b
0.0 c
74.5 d |
114.1 a
0.0 c
70.3 d |
117.0 a
0.0 c
69.9 d |
|
| B3LYP |
113.5 a
77.0 b
0.0 c
12.8 d |
126.6 a
221.8 b
0.0 c
45.7 d |
126.6 a
221.8 b
0.0 c
45.7 d |
117.4 a
248.0 b
0.0 c
61.6 d |
118.4 a
210.5 b
0.0 c
50.3 d |
118.1 a
221.8 b
0.0 c
61.3 d |
114.1 a
227.9 b
0.0 c
66.6 d |
121.6 a
225.0 b
0.0 c
54.6 d |
119.2 a
237.5 b
0.0 c
66.9 d |
115.6 a
221.0 b
0.0 c
60.0 d |
0.0 c
66.0 d |
114.4 a
235.5 b
0.0 c
66.7 d |
116.5 a
227.7 b
0.0 c |
115.4 a
232.9 b
0.0 c
66.1 d |
109.0 a
229.4 b
0.0 c
88.7 d |
112.3 a
231.2 b
0.0 c
66.8 d |
115.4 a
0.0 c
66.1 d |
|
| B3LYPultrafine |
|
|
|
|
118.4 a
0.0 c
50.3 d |
|
0.0 c
66.7 d |
|
|
|
|
|
|
115.4 a
0.0 c
66.1 d |
|
112.3 a
231.2 b
0.0 c
66.9 d |
|
|
| B3PW91 |
118.3 a
81.4 b
0.0 c
19.0 d |
131.2 a
223.2 b
0.0 c
54.1 d |
131.2 a
223.2 b
0.0 c
54.1 d |
121.0 a
248.7 b
0.0 c
68.4 d |
122.1 a
207.2 b
0.0 c
56.6 d |
121.6 a
218.8 b
0.0 c
67.9 d |
117.4 a
222.8 b
0.0 c
71.9 d |
123.8 a
217.2 b
0.0 c
59.2 d |
121.3 a
230.4 b
0.0 c
72.2 d |
118.8 a
216.9 b
0.0 c
66.6 d |
|
|
119.5 a
222.6 b
0.0 c
70.2 d |
117.5 a
225.2 b
0.0 c
71.4 d |
0.0 c
74.9 d |
114.7 a
0.0 c
71.9 d |
117.5 a
0.0 c
71.4 d |
|
| mPW1PW91 |
116.4 a
75.8 b
0.0 c
20.5 d |
126.9 a
216.2 b
0.0 c
55.3 d |
131.2 a
220.5 b
0.0 c
55.4 d |
120.9 a
247.4 b
0.0 c
70.2 d |
118.6 a
200.8 b
0.0 c
57.6 d |
118.2 a
212.7 b
0.0 c
69.1 d |
114.2 a
217.2 b
0.0 c
73.2 d |
120.2 a
210.2 b
0.0 c
60.0 d |
121.5 a
227.6 b
0.0 c
73.2 d |
119.2 a
214.6 b
0.0 c
67.8 d |
|
|
115.8 a
216.4 b
0.0 c
71.3 d |
114.0 a
218.8 b
0.0 c
72.5 d |
0.0 c
76.1 d |
|
114.0 a
0.0 c
72.5 d |
|
| M06-2X |
|
|
a
b
d |
|
a
b
d |
|
|
|
|
a
d |
|
|
|
a
d |
|
a
d |
|
|
| PBEPBE |
123.1 a
98.4 b
0.0 c
12.9 d |
129.0 a
223.9 b
0.0 c
47.5 d |
129.0 a
223.9 b
0.0 c
47.5 d |
121.3 a
248.8 b
0.0 c
62.0 d |
119.9 a
209.1 b
0.0 c
52.5 d |
119.4 a
220.3 b
0.0 c
63.4 d |
116.0 a
226.5 b
0.0 c
69.3 d |
123.2 a
221.5 b
0.0 c
56.7 d |
120.5 a
234.4 b
0.0 c
69.2 d |
116.3 a
217.4 b
0.0 c
62.3 d |
|
|
117.6 a
224.9 b
0.0 c
82.9 d |
116.3 a
228.7 b
0.0 c
68.9 d |
0.0 c
72.5 d |
113.3 a
0.0 c
70.0 d |
116.3 a
0.0 c
68.9 d |
|
| PBEPBEultrafine |
|
|
|
|
119.9 a
209.1 b
0.0 c
52.5 d |
|
|
|
|
|
|
|
|
|
|
|
|
|
| PBE1PBE |
|
|
|
|
123.0 a
204.4 b
0.0 c
58.5 d |
|
|
|
|
|
|
|
|
|
|
|
|
|
| HSEh1PBE |
|
222.2 b
0.0 c
55.2 d |
|
|
122.4 a
206.5 b
0.0 c
57.9 d |
|
223.0 b
0.0 c
73.6 d |
|
|
|
|
|
|
224.7 b
0.0 c
72.9 d |
|
|
|
|
| TPSSh |
|
|
|
|
118.8 a
191.3 b
0.0 c
43.0 d |
|
114.0 a
205.7 b
0.0 c
57.6 d |
|
|
115.2 a
199.1 b
0.0 c
52.0 d |
|
|
|
113.7 a
206.5 b
0.0 c
56.9 d |
|
|
|
|
| wB97X-D |
|
|
133.5 a
394.3 b
0.0 c
57.4 d |
|
a
b
d |
|
a
b
d |
|
a
b
d |
|
|
a
b
d |
a
b
d |
a
b
d |
|
a
b
d |
|
|
| B97D3 |
|
128.8 a
390.8 b
0.0 c
45.9 d |
|
|
118.5 a
380.1 b
0.0 c
49.7 d |
|
114.0 a
395.2 b
0.0 c
65.5 d |
|
118.4 a
402.4 b
0.0 c
65.8 d |
|
112.3 a
396.1 b
0.0 c
65.2 d |
114.0 a
401.7 b
0.0 c
66.0 d |
|
114.6 a
397.0 b
0.0 c
65.0 d |
|
111.7 a
395.5 b
0.0 c
66.0 d |
|
|
| |
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
cc-pV(T+d)Z |
daug-cc-pVTZ |
|---|
| Moller Plesset perturbation |
MP2 |
103.7 a
110.3 b
0.0 c
17.6 d |
129.2 a
243.9 b
0.0 c
49.0 d |
a
b
d |
120.0 a
277.4 b
0.0 c
69.4 d |
125.8 a
395.5 b
0.0 c
59.9 d |
124.4 a
233.2 b
0.0 c
67.8 d |
b |
130.2 a
400.9 b
0.0 c
62.0 d |
123.5 a
241.7 b
0.0 c
75.3 d |
124.3 a
230.7 b
0.0 c
69.2 d |
|
118.8 a
237.0 b
0.0 c
70.1 d |
123.7 a
243.5 b
0.0 c
73.4 d |
b |
113.4 a
0.0 c
75.7 d |
116.7 a
0.0 c
73.5 d |
119.7 a
0.0 c
73.1 d |
|
| MP2=FULL |
NC
NC |
129.2 a
243.9 b
0.0 c
49.0 d |
NC
NC |
NC
NC |
126.3 a
225.7 b
0.0 c
60.0 d |
125.0 a
233.4 b
0.0 c
67.8 d |
119.8 a
236.1 b
0.0 c
71.1 d |
130.7 a
232.0 b
0.0 c
62.8 d |
123.9 a
242.2 b
0.0 c
75.9 d |
127.2 a
233.6 b
0.0 c
72.1 d |
|
|
124.1 a
243.9 b
0.0 c
73.4 d |
120.5 a
233.4 b
0.0 c
73.6 d |
|
117.3 a
0.0 c
74.8 d |
NC |
|
| MP3 |
|
|
|
|
209.9 b
0.0 c
63.1 d |
|
0.0 c |
|
|
|
|
|
|
|
|
|
|
|
| MP3=FULL |
|
|
|
|
120.0 a
210.0 b
0.0 c
63.0 d |
|
114.0 a
218.7 b
0.0 c
72.5 d |
|
|
|
|
|
|
|
|
|
|
|
| MP4 |
|
NC
NC |
|
|
122.1 a
223.0 b
0.0 c
55.7 d |
|
|
|
NC |
|
|
|
|
|
|
|
|
|
| B2PLYP |
|
|
|
|
121.7 a
217.6 b
0.0 c
53.0 d |
|
|
|
|
119.2 a
0.0 c
62.7 d |
|
|
|
117.4 a
234.5 b
0.0 c
67.8 d |
|
|
|
|
| Configuration interaction |
CID |
|
NC
NC |
NC
NC |
NC
NC |
123.1 a
210.9 b
0.0 c
85.9 d |
|
|
NC
NC |
|
|
|
|
|
|
|
|
|
|
| CISD |
|
NC
NC |
NC
NC |
NC
NC |
123.5 a
211.8 b
0.0 c
84.9 d |
|
|
NC
NC |
|
|
|
|
|
|
|
|
|
|
| |
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
cc-pV(T+d)Z |
daug-cc-pVTZ |
|---|
| Quadratic configuration interaction |
QCISD |
NC |
120.2 a
222.7 b
0.0 c
71.4 d |
NC
NC |
NC
NC |
119.6 a
216.6 b
0.0 c
58.0 d |
117.8 a
222.3 b
0.0 c
64.8 d |
113.0 a
224.4 b
0.0 c
67.8 d |
123.5 a
221.6 b
0.0 c
59.9 d |
116.7 a
0.0 c
71.0 d |
118.8 a
0.0 c
66.3 d |
|
|
NC
NC |
114.1 a
0.0 c
69.9 d |
|
111.4 a
0.0 c
70.1 d |
|
|
| QCISD(T) |
|
|
|
|
NC
NC |
|
NC |
|
NC |
|
|
|
|
|
|
|
|
|
| Coupled Cluster |
CCD |
NC |
NC
NC |
NC
NC |
NC
NC |
118.5 a
211.0 b
0.0 c
60.9 d |
NC
NC |
NC
NC |
NC
NC |
NC |
NC |
|
|
NC
NC |
|
|
|
|
|
| CCSD |
|
|
|
|
119.3 a
214.4 b
0.0 c
58.9 d |
|
NC |
|
NC |
118.5 a
0.0 c
67.1 d |
|
|
|
113.9 a
0.0 c
70.5 d |
|
|
|
|
| CCSD=FULL |
|
|
|
|
119.8 a
0.0 c
58.9 d |
|
|
|
|
121.7 a
0.0 c
70.0 d |
|
|
|
114.9 a
0.0 c
70.8 d |
|
|
|
|
| CCSD(T) |
|
|
|
|
NC
NC |
|
NC |
|
NC |
|
|
|
NC |
NC |
NC |
|
|
|
| |
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
cc-pV(T+d)Z |
daug-cc-pVTZ |
|---|