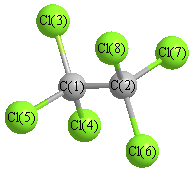
Vibrational Frequencies calculated at MP2=FULL/aug-cc-pVTZ
| Mode Number |
Symmetry |
Frequency
(cm-1) |
Scaled Frequency
(cm-1) |
IR Intensities
(km mol-1) |
Raman Act
(Å4/u) |
Dep P |
Dep U |
|---|
| 1 |
A1g |
1031 |
981 |
0.00 |
3.03 |
0.34 |
0.50 |
| 2 |
A1g |
451 |
429 |
0.00 |
25.27 |
0.02 |
0.03 |
| 3 |
A1g |
229 |
218 |
0.00 |
1.01 |
0.32 |
0.48 |
| 4 |
A1u |
89 |
85 |
0.00 |
0.00 |
0.00 |
0.00 |
| 5 |
A2u |
707 |
672 |
38.36 |
0.00 |
0.00 |
0.00 |
| 6 |
A2u |
389 |
370 |
0.55 |
0.00 |
0.00 |
0.00 |
| 7 |
Eg |
893 |
849 |
0.00 |
11.62 |
0.75 |
0.86 |
| 7 |
Eg |
893 |
849 |
0.00 |
11.62 |
0.75 |
0.86 |
| 8 |
Eg |
354 |
337 |
0.00 |
4.48 |
0.75 |
0.86 |
| 8 |
Eg |
354 |
337 |
0.00 |
4.48 |
0.75 |
0.86 |
| 9 |
Eg |
231 |
220 |
0.00 |
2.10 |
0.75 |
0.86 |
| 9 |
Eg |
231 |
220 |
0.00 |
2.10 |
0.75 |
0.86 |
| 10 |
Eu |
827 |
786 |
168.98 |
0.00 |
0.00 |
0.00 |
| 10 |
Eu |
827 |
786 |
168.98 |
0.00 |
0.65 |
0.00 |
| 11 |
Eu |
283 |
270 |
0.00 |
0.00 |
0.00 |
0.00 |
| 11 |
Eu |
283 |
270 |
0.00 |
0.00 |
0.00 |
0.00 |
| 12 |
Eu |
171 |
163 |
0.06 |
0.00 |
0.00 |
0.00 |
| 12 |
Eu |
171 |
163 |
0.06 |
0.00 |
0.00 |
0.00 |
Unscaled Zero Point Vibrational Energy (zpe) 4206.5 cm
-1
Scaled (by 0.9514) Zero Point Vibrational Energy (zpe) 4002.1 cm
-1
See section
III.C.1 List or set vibrational scaling factors
to change the scale factors used here.
See section
III.C.2
Calculate a vibrational scaling factor for a given set of molecules
to determine the least squares best scaling factor.