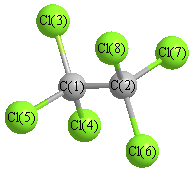
Vibrational Frequencies calculated at MP2/Def2TZVPP
| Mode Number |
Symmetry |
Frequency
(cm-1) |
Scaled Frequency
(cm-1) |
IR Intensities
(km mol-1) |
Raman Act
(Å4/u) |
Dep P |
Dep U |
|---|
| 1 |
A1g |
994 |
946 |
0.00 |
2.83 |
0.41 |
0.58 |
| 2 |
A1g |
446 |
424 |
0.00 |
18.54 |
0.03 |
0.06 |
| 3 |
A1g |
226 |
215 |
0.00 |
0.93 |
0.54 |
0.70 |
| 4 |
A1u |
88 |
84 |
0.00 |
0.00 |
0.00 |
0.00 |
| 5 |
A2u |
697 |
664 |
44.17 |
0.00 |
0.00 |
0.00 |
| 6 |
A2u |
386 |
367 |
0.67 |
0.00 |
0.00 |
0.00 |
| 7 |
Eg |
878 |
836 |
0.00 |
9.52 |
0.75 |
0.86 |
| 7 |
Eg |
878 |
836 |
0.00 |
9.52 |
0.75 |
0.86 |
| 8 |
Eg |
348 |
332 |
0.00 |
5.76 |
0.75 |
0.86 |
| 8 |
Eg |
348 |
332 |
0.00 |
5.76 |
0.75 |
0.86 |
| 9 |
Eg |
227 |
216 |
0.00 |
3.06 |
0.75 |
0.86 |
| 9 |
Eg |
227 |
216 |
0.00 |
3.06 |
0.75 |
0.86 |
| 10 |
Eu |
815 |
776 |
182.44 |
0.00 |
0.31 |
0.00 |
| 10 |
Eu |
815 |
776 |
182.44 |
0.00 |
0.00 |
0.00 |
| 11 |
Eu |
282 |
268 |
0.00 |
0.00 |
0.00 |
0.00 |
| 11 |
Eu |
282 |
268 |
0.00 |
0.00 |
0.00 |
0.00 |
| 12 |
Eu |
168 |
159 |
0.10 |
0.00 |
0.00 |
0.00 |
| 12 |
Eu |
168 |
159 |
0.10 |
0.00 |
0.00 |
0.00 |
Unscaled Zero Point Vibrational Energy (zpe) 4136.6 cm
-1
Scaled (by 0.9515) Zero Point Vibrational Energy (zpe) 3936.0 cm
-1
See section
III.C.1 List or set vibrational scaling factors
to change the scale factors used here.
See section
III.C.2
Calculate a vibrational scaling factor for a given set of molecules
to determine the least squares best scaling factor.