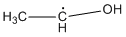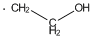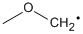Methods with standard basis sets
|
|
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
cc-pVQZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
aug-cc-pVQZ |
daug-cc-pVTZ |
| hartree fock |
HF |
0.0 a
102.9 b
140.9 c
115.9 d |
0.0 a
42.2 b
65.3 c
75.5 d |
0.0 a
42.2 b
65.3 c
75.5 d |
0.0 a
32.4 b
52.6 c
89.9 d |
0.0 a
26.1 b
49.0 c
65.3 d |
0.0 a
12.4 b
35.2 c
63.9 d |
0.0 a
6.0 b
26.4 c
59.5 d |
0.0 a
23.8 b
45.8 c
62.4 d |
0.0 a
8.0 b
29.8 c
59.9 d |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC
NC |
NC
NC
NC |
|
0.0 a
4.9 b
26.4 c
55.6 d |
| ROHF |
|
0.0 a
50.7 b
65.2 c
75.5 d |
0.0 a
50.7 b
65.2 c
75.5 d |
0.0 a
41.4 b
52.6 c
89.9 d |
0.0 a
36.5 b
48.9 c
65.3 d |
0.0 a
22.9 b
35.3 c
63.9 d |
0.0 a
30.6 b
40.9 c
61.3 d |
0.0 a
34.4 b
45.9 c
62.4 d |
0.0 a
18.8 b
29.8 c
59.9 d |
|
|
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC
NC |
NC
NC
NC |
|
|
| density functional |
LSDA |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
0.0 a
-21.2 b
31.4 c
32.3 d |
0.0 a
-28.5 b
23.2 c
13.6 d |
0.0 a
-39.3 b
12.4 c
13.6 d |
0.0 a
-43.8 b
2.3 c
14.5 d |
0.0 a
-30.1 b
21.3 c
16.0 d |
0.0 a
-45.4 b
5.3 c
13.4 d |
0.0 a
-43.7 b
6.9 c
7.0 d |
|
NC
NC |
0.0 a
-43.7 b
7.9 c
9.4 d |
0.0 a
-49.6 b
-1.7 c
9.0 d |
|
0.0 a
-49.5 b
-5.4 c
12.9 d |
0.0 a
-51.6 b
-5.5 c
8.8 d |
|
|
| BLYP |
0.0 a
27.7 b
99.9 c
26.6 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-0.6 b
41.0 c
39.2 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-26.8 b
9.9 c
21.5 d |
0.0 a
-14.1 b
28.5 c
21.9 d |
0.0 a
-27.9 b
14.3 c
19.9 d |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
0.0 a
-31.6 b
7.8 c
16.4 d |
|
|
|
|
|
| B1B95 |
0.0 a
41.0 b
113.7 c
51.1 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-0.6 b
40.7 c
52.4 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-28.2 b
9.8 c
28.0 d |
0.0 a
-12.7 b
29.2 c
29.9 d |
0.0 a
-28.1 b
13.3 c
27.4 d |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
0.0 a
-32.4 b
4.0 c
28.1 d |
0.0 a
-34.2 b
3.7 c
21.6 d |
|
|
| B3LYP |
0.0 a
42.1 b
108.9 c
46.0 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
1.1 b
39.5 c
47.5 d |
0.0 a
-6.9 b
32.1 c
28.8 d |
NC
NC
NC |
0.0 a
-26.7 b
7.8 c
25.0 d |
0.0 a
-12.6 b
26.6 c
26.3 d |
0.0 a
-27.2 b
11.6 c
24.0 d |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
0.0 a
-31.1 b
5.6 c
19.7 d |
0.0 a
-33.1 b |
0.0 a
-32.4 b
0.7 c
24.1 d |
0.0 a
-34.0 b
1.2 c
18.7 d |
0.0 a
-34.6 b |
|
| B3LYPultrafine |
|
|
|
|
NC
NC
NC |
|
|
|
|
|
|
|
|
|
|
|
0.0 a
-33.9 b
1.2 c
18.7 d |
|
|
| B3PW91 |
0.0 a
43.6 b
110.7 c
53.6 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-2.0 b
36.9 c
51.3 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-30.4 b
5.1 c
26.7 d |
0.0 a
-15.0 b
24.1 c
28.8 d |
0.0 a
-30.5 b
8.3 c
26.3 d |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
0.0 a
-34.6 b
2.5 c
21.7 d |
|
|
|
|
|
| mPW1PW91 |
0.0 a
47.8 b
112.9 c
59.0 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-0.4 b
37.2 c
54.3 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-29.7 b
4.9 c
28.4 d |
0.0 a
-13.4 b
24.6 c
30.8 d |
0.0 a
-29.2 b
8.6 c
28.2 d |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
0.0 a
-33.3 b
3.0 c
23.6 d |
|
0.0 a
-33.2 b
-0.1 c
28.5 d |
0.0 a
-35.8 b
-0.7 c
22.6 d |
|
|
| M06-2X |
0.0 a
41.3 b
46.6 d |
NC
NC
NC |
0.0 a
5.9 b
45.3 c
35.1 d |
0.0 a
-2.4 b
33.5 c
48.3 d |
0.0 a
-16.7 b
20.3 c
20.6 d |
0.0 a
-28.4 b
8.7 c
19.8 d |
0.0 a
-34.6 b
-1.5 c
17.7 d |
0.0 a
-18.9 b
17.0 c
19.7 d |
0.0 a
-34.3 b
1.1 c
17.0 d |
0.0 a
-34.7 b
3.0 c |
|
NC
NC |
0.0 a
-34.4 b
3.8 c
15.1 d |
0.0 a
-39.4 b
-5.4 c
12.0 d |
|
0.0 a
-40.8 b
-8.5 c |
0.0 a
-41.4 b
-8.4 c
11.3 d |
|
|
| PBEPBE |
0.0 a
27.3 b
102.5 c
33.8 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-4.3 b
39.2 c
44.6 d |
NC
NC
NC |
0.0 a
-25.2 b
18.6 c
25.4 d |
0.0 a
-31.3 b
7.6 c
24.6 d |
0.0 a
-16.8 b
27.0 c
25.8 d |
0.0 a
-31.8 b
11.6 c
23.5 d |
0.0 a
-29.5 b
13.6 c
19.3 d |
|
NC
NC |
0.0 a
-29.0 b
15.3 c
21.9 d |
0.0 a
-36.1 b
4.9 c
19.4 d |
|
0.0 a
-36.0 b
1.4 c
24.2 d |
0.0 a
-38.6 b
0.5 c
18.8 d |
|
|
| HSEh1PBE |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
0.0 a
0.4 b
38.4 c
55.4 d |
NC
NC
NC |
NC
NC
NC |
|
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC
NC |
NC
NC
NC |
|
|
| TPSSh |
|
|
|
|
0.0 a
-2.3 b
36.0 c
28.4 d |
|
NC
NC
NC |
|
|
NC
NC
NC |
|
|
|
NC
NC
NC |
|
|
|
|
|
| wB97X-D |
|
|
0.0 a
0.5 b
45.4 c
38.4 d |
|
0.0 a
-13.3 b
25.9 c
29.0 d |
|
0.0 a
-31.1 b
4.2 c
26.0 d |
|
0.0 a
-30.4 b
8.3 c
26.0 d |
|
|
0.0 a
-30.6 b
6.4 c
26.9 d |
0.0 a
-14.8 b
4.2 c
26.0 d |
0.0 a
-33.5 b
3.7 c
21.7 d |
|
|
0.0 a
-35.5 b
0.5 c
21.1 d |
|
|
| B97D3 |
|
NC
NC |
|
|
NC
NC |
|
0.0 a
6.0 c
23.3 d |
|
0.0 a
10.2 c
23.1 d |
|
0.0 a
-36.7 b
0.2 c
17.0 d |
0.0 a
7.0 c
23.7 d |
|
0.0 a
3.9 c
18.5 d |
|
|
0.0 a
-0.4 c
17.9 d |
|
|
| |
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
cc-pVQZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
aug-cc-pVQZ |
daug-cc-pVTZ |
|---|
| Moller Plesset perturbation |
MP2 |
0.0 a
79.6 b
128.1 c
87.0 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-13.2 b
15.4 c
42.3 d |
0.0 a
-27.8 b
5.6 c
17.4 d |
NC
NC
NC |
NC
NC |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
0.0 a
-54.6 b
-22.8 c
2.9 d |
|
NC
NC
NC |
NC
NC |
|
|
| MP2=FULL |
0.0 a
79.7 b
128.3 c
87.3 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-13.3 b
15.3 c
42.1 d |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC |
NC
NC
NC |
NC
NC
NC |
|
NC
NC
NC |
NC
NC
NC |
|
|
| ROMP2 |
0.0 a
80.0 b
130.6 c
87.4 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-13.9 b
18.6 c
41.8 d |
NC
NC |
NC
NC |
NC
NC |
NC
NC |
NC
NC |
NC
NC |
|
|
NC
NC |
NC
NC |
|
NC
NC |
|
|
|
| MP3 |
|
|
|
|
NC
NC
NC |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| MP3=FULL |
|
|
|
|
NC
NC
NC |
|
NC
NC
NC |
|
|
|
|
|
|
|
|
|
|
|
|
| MP4 |
|
NC
NC |
|
|
NC
NC |
|
|
|
NC
NC |
|
|
|
|
|
|
|
|
|
|
| Configuration interaction |
CID |
|
NC
NC
NC |
NC
NC
NC |
0.0 a
6.5 b
31.3 c
62.6 d |
NC
NC
NC |
|
|
0.0 a
-5.8 b
23.1 c
37.5 d |
|
|
|
|
|
|
|
|
|
|
|
| CISD |
|
NC
NC
NC |
NC
NC
NC |
0.0 a
6.5 b
32.5 c
61.3 d |
NC
NC
NC |
|
|
0.0 a
-5.4 b
24.6 c
37.2 d |
|
|
|
|
|
|
|
|
|
|
|
| |
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
cc-pVQZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
aug-cc-pVQZ |
daug-cc-pVTZ |
|---|
| Quadratic configuration interaction |
QCISD |
|
NC
NC
NC |
NC
NC
NC |
0.0 a
-1.9 b
27.1 c
50.4 d |
NC
NC
NC |
0.0 a
-36.0 b
-3.1 c
14.5 d |
NC
NC
NC |
NC
NC
NC |
0.0 a
-43.5 b
-10.6 c
11.8 d |
NC
NC |
|
NC
NC |
NC
NC
NC |
NC
NC |
|
NC
NC |
|
|
|
| QCISD(T) |
|
|
|
|
NC
NC |
|
|
|
|
|
|
|
0.0 a
-31.6 b
19.8 d |
|
|
|
|
|
|
| Coupled Cluster |
CCD |
|
NC
NC
NC |
NC
NC
NC |
0.0 a
2.0 b
27.3 c
57.5 d |
NC
NC
NC |
NC
NC
NC |
NC
NC
NC |
0.0 a
-10.8 b
19.5 c
34.0 d |
NC
NC
NC |
NC
NC |
|
NC
NC |
NC
NC
NC |
NC
NC |
|
NC
NC |
|
|
|
| CCSD |
|
|
|
|
NC
NC
NC |
|
|
|
|
|
|
|
NC
NC
NC |
0.0 a
-29.9 b
24.3 d |
|
|
|
|
|
| CCSD(T) |
|
|
|
|
NC
NC |
|
|
|
|
|
|
|
NC
NC |
|
|
NC
NC |
|
|
|
| CCSD(T)=FULL |
|
|
|
|
NC
NC |
|
|
|
|
|
|
|
NC
NC |
|
|
NC
NC |
|
|
|
| |
STO-3G |
3-21G |
3-21G* |
6-31G |
6-31G* |
6-31G** |
6-31+G** |
6-311G* |
6-311G** |
6-31G(2df,p) |
6-311+G(3df,2p) |
TZVP |
cc-pVDZ |
cc-pVTZ |
cc-pVQZ |
aug-cc-pVDZ |
aug-cc-pVTZ |
aug-cc-pVQZ |
daug-cc-pVTZ |
|---|